Expanding the Galaxy Platform
Galaxy is an open-source, web-accessible, integrated informatics solution for data-intensive research in biological and medical science. It is designed so that biologists without programming experience can use it, and it captures information so that computational analyses can be duplicated by other users. It also allows users to share, collaborate, and publish their analyses via the web.
The Galaxy framework was originally designed at Penn State as a web interface for bioinformatics analysis of genomic data, especially Next Generation Sequencing (NGS) data. It is a web server that allows users to specify tools – a tool is a command-line application with its parameter options - to run via web forms. Files used as inputs for tools and those that are generated are labeled with file formats (datatypes) that allow tools to be connected together as workflows.
Penn State’s original Galaxy framework was created with the intention that it could be extended and adapted to any field that uses command-line applications to process and analyze data. This can be done by adding tools that represent applications needed for analysis and defining file formats as Galaxy datatypes.
In a project called Galaxy-P, Professor Tim Griffin (Biochemistry, Molecular Biology, and Biophysics; Director, Center for Mass Spectrometry and Proteomics (CMSP)) collaborated with MSI’s Research Informatics Solutions (RIS) group to extend the set of Galaxy tools and datatypes to perform proteomics analysis of data generated by mass spectrometry. (Proteomics is the study of the proteome, the entire set of proteins in an organism or system.) In a three-year project, the CMSP (led by Managing Director Pratik Jagtap) and MSI staff worked on tools to analyze proteomics data gathered from mass spectrometry. These included analysis tools, visualizers, and file-conversion tools. University of Minnesota staff worked with Penn State to improve and extend the Galaxy framework to handle the requirements of proteomics.
The U team is also promoting this work through publications in journals and oral presentations at conferences, in order to inform the larger proteomics researcher community about these new tools. Over the course of the project, at least 11 presentations were given at various international conferences to describe the capabilities and architecture of the solution. Galaxy-P has also been featured in professional journals; some examples are (names of MSI Principal Investigators are in bold):
Boekel, Jorrit, John M. Chilton, Ira R. Cooke, Peter L. Horvatovich, Pratik D. Jagtap, Lukas Kall, Janne Lehtio, Pieter Lukasse, Perry D. Moerland, and Timothy J. Griffin. 2015. Multi-omic data analysis using Galaxy. Nature Biotechnology 33(2)(JAN 01): 137-9.
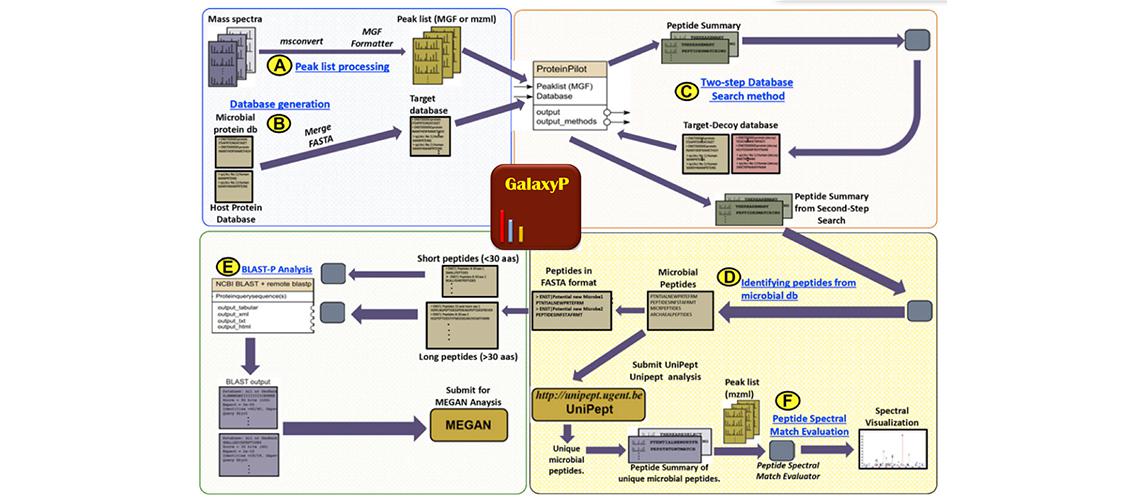
Jagtap, Pratik D., Alan Blakely, Kevin Murray, Shaun Stewart, Joel Kooren, James E. Johnson, Nelson L. Rhodus, Joel D. Rudney, and Timothy J. Griffin. 2015. Metaproteomic analysis using the Galaxy framework. Proteomics. DOI: 10.1002/pmic.201500074.
Jagtap, Pratik, James E. Johnson, Getiria Onsongo, Frederik Sadler , Kevin Murray, Y Wang, Gloria Shenykman, Sricharan Bandhakavi, Lloyd M Smith and Timothy J. Griffin. 2014. Flexible and accessible workflows for improved proteogenomic analysis using the Galaxy framework. Journal of Proteome Research 13(12): 5898-908.
The U has recently received an NSF grant for the continued expansion of Galaxy. This project will develop tools for “multi-omic” analysis, which will leverage Galaxy’s ability to connect software for different fields into sophisticated workflows. Galaxy-M, which is one part of this project, will add additional tools for metabolomics analysis. (Metabolomics is the study of processes involving metabolites, the products of metabolism.) Associate Professor Adrian Hegeman (Horticultural Science), who is a co-investigator on the NSF grant, did initial work on Galaxy-M through a fellowship from the University of Minnesota Informatics Institute. This work was featured in a story on the Department of Horticultural Science website in July 2015.