Restoring Neural Connections in Hurler Syndrome using PS Gene-Editing System
Hurler syndrome, a genetic brain disorder also known as mucopolysaccharidosis type I (MPS I), causes severe cognitive and physical abnormalities and early death. Current treatments for this disorder are inadequate. Recent research by MSI PIs has shown a possible new method of treatment for this disease and other disorders. Using a method developed at the U of M called the PS gene-editing system, the researchers were able to restore disrupted neural networks in the brains of mice affected with Hurler syndrome. The researchers used resting-state function MRI imaging to examine the brains pre- and post-treatment. The research was published recently in the journal Scientific Reports: W. Zhu, L. Ou, L. Zhang, I.H. Clark, Y. Zhang, X.-H. Zhu, C.B. Whitley, P.B. Hackett, W.C. Low, W. Chen. Mapping brain networks in MPS I Mice and Their Restoration Following Gene Therapy. Scientific Reports 13: 12716 (2023). doi: 10.1038/s41598-023-39939-0. A story about this paper appears on the U of M News site: PS gene-editing shown to restore neural connections lost in brain disorder.
The MSI PIs who are co-authors on this paper are Professor Walter Low (Neurosurgery) and Associate Professor Lin Zhang (Biostatistics). Professor Low uses MSI for a variety of projects that study the brain and developmental biology. Professor Zhang uses MSI for several projects on Bayesian analysis of high-dimensional genomics and image data. Co-author Dr. Ying Zhang is an Informatics Analyst at MSI.
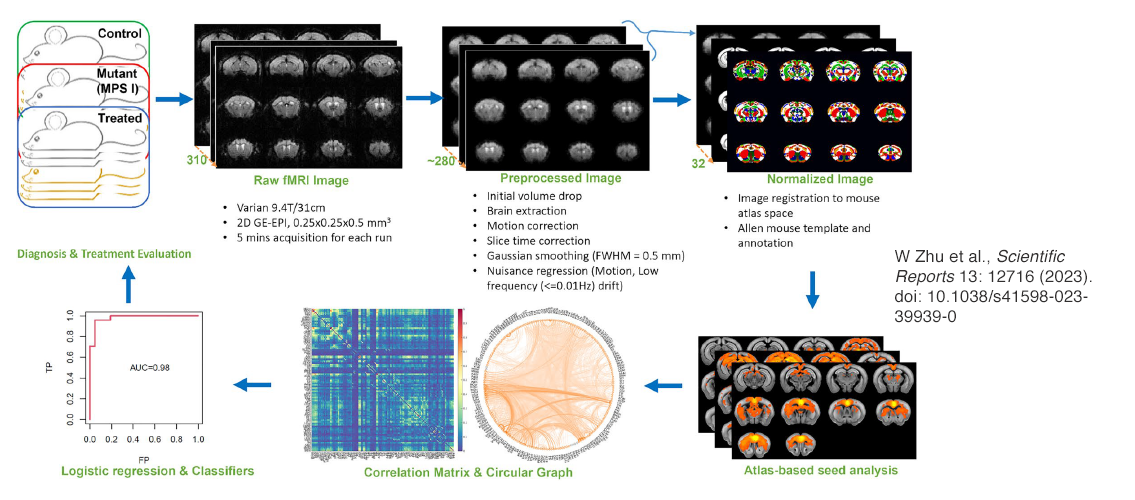
Image description: Resting-state functional magnetic resonance imaging data analysis pipeline. Three cohorts of mice were scanned in the 9.4 Tesla Varian MRI scanner with the bore size of 31 cm. FMRI raw data were acquired using the 2D gradient echo-based echo planer imaging (GE-EPI) sequence with a nominal spatial resolution of 0.25 × 0.25 × 0.5 mm3 and temporal resolution of 1 s. Three to five fMRI runs were acquired for each mouse with 5 min acquisition for each run. Raw fMRI images were preprocessed using the standard fMRI preprocessing pipeline and thereafter normalized to Allen brain mouse atlas. By using labeled brain regions as seedings, Pearson’s correlations were calculated between the seeding region and all fMRI voxels to generate RSNs. The region-to-region correlations were also calculated to derive the correlation matrix, which can be visualized using circular graph. Logistic regression was then conducted on either the correlation matrices or the RSNs for each cohort to generate spatial and ROI (region of interest) classifiers that were used to predict the three cohorts. Image and description. W. Zhu et al., Scientific Reports 13: 12716 (2023). doi: 10.1038/s41598-023-39939-0.